The 3+ days at useR!2019 in Toulouse were packed with great talks1 and good food - hence the amuse-bouches word play.
Here are some R code bits from the conference. Hopefully convincing enough to start using a new package or change a workflow. Not everything was brand-new, but it was helpful to have someone talking through their inspiration and examples.
Check out the speakers’ materials - soon there will be recordings too. Some of the examples are also copied straight from the speakers’ slide decks.
1. tidy eval
Speaker: Lionel Henry (Slides)
I never warmed up to the bang-bangs and enquo’s. Hence the new and more straight forward {{ }} (read: curly curly) for functional programming {{ arg }} in the tidyverse feel like a game-changer.
For those more familiar with the previous framework: {{ arg }} is a shortcut for !!enquo(arg).
dplyr example
Let’s say you have a dataset, here iris, and you want to compute the average Petal.Length for each Species:
library(dplyr)##
## Attaching package: 'dplyr'## The following objects are masked from 'package:stats':
##
## filter, lag## The following objects are masked from 'package:base':
##
## intersect, setdiff, setequal, unioniris %>%
group_by(Species) %>%
summarise(avg = mean(Petal.Length, na.rm = TRUE))## # A tibble: 3 x 2
## Species avg
## <fct> <dbl>
## 1 setosa 1.46
## 2 versicolor 4.26
## 3 virginica 5.55You can use the “curly-curly” brackets if you want to turn this small bit of code into a function group_mean() with a data, by and var argument2 (and want to pass the variables on in an unquoted way):
group_mean <- function(data, by, var) {
data %>%
group_by({{ by }}) %>%
summarise(avg = mean({{ var }}, na.rm = TRUE))
}We can then apply group_mean() to any dataset that has a grouping and a continuous variable, for example, the mammals sleep dataset in ggplot2:
library(ggplot2)
group_mean(data = msleep, by = vore, var = sleep_total)ggplot2 example
Another common tidy eval application is ggplot2. In the example below, we want a customised plot: a scatterplot with a geom_smooth on top of it.
library(ggplot2)
theme_set(theme_bw())
ggplot(data = iris,
aes(x = Sepal.Length, y = Petal.Length, group = Species, color = Species)
) +
geom_point() +
geom_smooth(method = "lm") +
ggtitle("Pack this plot into a function.")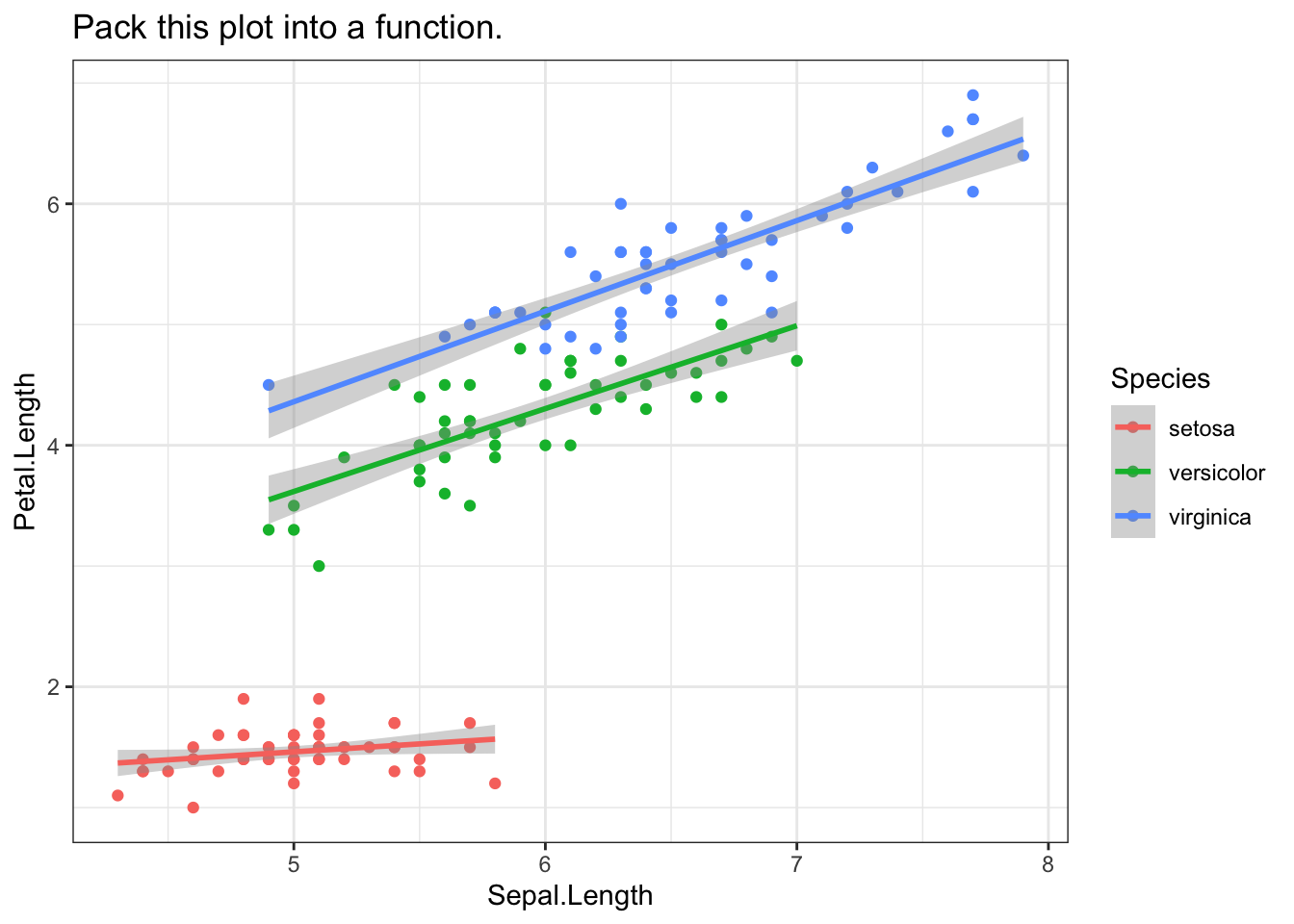
Again, we can wrap the “curly-curly” brackets around the arguments and apply them to a different dataset.
plot_point_smooth <- function(data, x, y, gr = NULL, method = "lm") {
ggplot(data = data,
aes({{ x }}, {{ y }}, group = {{ gr }}, color = {{ gr }})
) +
geom_point() +
geom_smooth(method = method)
}plot_point_smooth(msleep, x = sleep_total, y = sleep_rem, gr = NULL) +
ggtitle("Tidy eval with the msleep dataset")2. usethis
Speaker: Jenny Bryan (Slides + Material + Demo)
# install.packages("usethis")
library(usethis)Once upon a time, there was the package devtools. Then devtools became too large, and now the usethis package is taking over some of the convenience functions for workflows.
usethis is all about avoiding to copy+pasting. For example, there is a function to edit the .Rprofile called usethis::edit_r_profile(). Whenever there is a slightly complicated task ahead (say restarting R), the usethis package will talk you through the whole process.
There are lots of use_* to add or modify something to/in a project/package and three functions to create a package, a project or a github fork:
create_package()create_project()create_from_github()
Create a package
If you want to create a package, do the following (see also screencast):
## 1. create the package skeleton
create_package("~/tmp/mypackage")
## 2. use git
use_git()
## 3. add a license
use_mit_license()
## 4. run check
# install.packages("devtools")
devtools::check()
## 5. commit all files with git
## 6. set up git + github
use_github()
## will update the DESCRIPTION file
## 7. install the package
devtools::install()
## 8. add a rmarkdown readme file
use_readme_rmd()
## knit + commit + push
## 9. clean up if this was only a demo
## install.packages("fs")
## fs::dir_delete("~/tmp/mypackage")3. pak
Speaker: Gábor Csárdi (Slides)
# install.packages("pak") ## or
# devtools::install_github("r-lib/pak")It seems like pak will make package installation - conventional and for projects - more intuitive. Before installing anything, pak will give you a heads up on what will be installed or if there are any conflicts.
pak has two main functions: pak::pkg_* and pak:::proj_*
Conventional package installation
Play around with usethis3 and see what happens:
pak::pkg_install("usethis")
pak::pkg_remove("usethis")
pak::pkg_install("r-lib/usethis")
pak::pkg_status("usethis")Package installation for projects
First, create a project with usethis, then install R packages directly into the project.
usethis::create_project("~/tmp/test")
## check the directory
dir()
## initialise a dedicated R packages folder
pak:::proj_create()
## check the directory again
dir()
## check the DESCRIPTION file
readLines("DESCRIPTION")
## install usethis
pak:::proj_install("usethis")
## this installs dependencies into a private project library
readLines("DESCRIPTION")
## remove the project folder again
fs::dir_delete("~/tmp/test")4. Reshaping data
Speaker: Hadley Wickham (Demo)
# install.packages("tidyverse/tidyr")
library(tidyr)What a history reshaping data in R already has! From reshape to melt + cast, over to gather + spread and now pivot_long + pivot_wide. Reshaping data stays a mind-bending task, but hopefully, these pivot_* functions will make life easier.
# devtools::install_github("chrk623/dataAnim")
# Master's Thesis project by Charco Hui
library(dataAnim)
## Our two toy datasets
datoy_wide## Name English Maths
## 1 Ben 19.0 58.5
## 2 Sam 6.7 51.8
## 3 Sarah 14.9 45.1datoy_long## Name Subject Score
## 1 Ben Maths 10.0
## 2 Ben English 63.7
## 3 Sam Maths 52.9
## 4 Sam English 75.6
## 5 Alex Maths 88.8
## 6 Alex English 92.2Let’s reshape the datasets4:
## lets make it longer
datoy_wide %>%
pivot_longer(-Name, names_to = "Subject", values_to = "Score")
## lets make it wider
datoy_long %>%
dplyr::mutate(Time = 1:nrow(datoy_long)) %>%
pivot_wider(names_from = "Subject", values_from = c("Score", "Time"))5. vroom
Speaker: Jim Hester (Slides + Screencast)
## install.packages("vroom")
library(vroom)Importing large datasets into R can be a painful task. Especially if you only need a subset of the columns. And apparently, our thoughts drift off after 10 sec5 staring at the screen where it is still loading the dataset.
data.table::fread() is always here to help. But now comes vroom!
Get some large’ish data
First, we need some large dataset. To not burden our laptops too much6, we will go for some exome based GWAS results.
## Source: https://portals.broadinstitute.org/collaboration/giant/index.php/GIANT_consortium_data_files
path_to_file_1 <- "Height_AA_add_SV.txt.gz"
path_to_file_2 <- "BMI_African_American.fmt.gzip"
## Height
download.file(
"https://portals.broadinstitute.org/collaboration/giant/images/8/80/Height_AA_add_SV.txt.gz",
path_to_file_1)
## BMI
download.file(
"https://portals.broadinstitute.org/collaboration/giant/images/3/33/BMI_African_American.fmt.gzip",
path_to_file_2)
## File size
## install.packages("fs")
fs::file_size(path_to_file_1)## 4.39Mfs::file_size(path_to_file_2)## 13.3MThe two datasets have a mix of characters, numbers and decimals7.
vroom vs DT
Here is how vroom works and a basic comparison to data.table::fread (I let you do the proper benchmarking yourself).
library(dplyr)
## With vroom
giant_vroom <- vroom::vroom(path_to_file_1)
giant_vroom_subset <- giant_vroom %>% select(CHR, POS) %>% filter(CHR == 1)
## The equivalent with data.table
giant_DT <- data.table::fread(path_to_file_1)
giant_DT_subset <- giant_DT %>% select(CHR, POS) %>% filter(CHR == 1)col_select for the win
## Selecting columns
giant_vroom_select <- vroom::vroom(path_to_file_1,
col_select = list(SNPNAME, ends_with("_MAF")))
head(giant_vroom_select)
## Preventing columns from being imported
giant_vroom_remove <- vroom::vroom(path_to_file_1,
col_select = -ExAC_AFR_MAF)
head(giant_vroom_remove)
## Renaming on the fly
giant_vroom_rename <- vroom::vroom(path_to_file_1,
col_select = list(p = Pvalue, everything()))
head(giant_vroom_rename)Combining multiple datasets
data_combined <- vroom::vroom(
c(path_to_file_1, path_to_file_2),
id = "path")
table(data_combined$path)6. data.table
Speaker: Arun Srinivasan (Slides)
data.table has a pretty cool feature8:
# install.packages("data.table")
library(data.table)## Warning: package 'data.table' was built under R version 3.5.2##
## Attaching package: 'data.table'## The following object is masked from 'package:dataAnim':
##
## :=## The following objects are masked from 'package:dplyr':
##
## between, first, last## Create a giant data.table
p <- 2e6
dat <- data.table(x = sample(1e5, p, TRUE), y = runif(p))
## Let's select a few rows
system.time(
tmp <- dat[x %in% 2000:3000 ]
)
## do the same operation again
system.time(
tmp <- dat[x %in% 2000:3000 ]
)7. rray
Speaker: Davis Vaughan (Slides)
# devtools::install_github("r-lib/rray")
## may take some timerray can do two things that are otherwise annoying/counter-intuitive in R:
- broadcasting (recycling dimensions)
- subsetting (
bag[,1, drop = FALSE])
Matrices with base-r
Let’s look at an example of matrix operations in base-r9.
First, we want to add two matrices with similar dimensions:
mat_1 <- matrix(c(15, 10, 8, 6, 12, 9), byrow = FALSE, nrow = 2)
mat_2 <- matrix(c(5, 2, 3), nrow = 1)
## broadcasting won't work ❌
mat_1 + mat_2## Error in mat_1 + mat_2: non-conformable arraysNext, we want to select one matrix column:
dim(mat_1[,2:3]) ## selecting two columns is fine## [1] 2 2## subsetting won't preserve the matrix class ❌
dim(mat_1[,1]) ## why not 2x1?## NULLlength(mat_1[,1]) ## ah, it turned into a vector!## [1] 2dim(mat_1[,1, drop = FALSE]) ## but with drop = FALSE we can keep it a matrix## [1] 2 1Matrices with rray
Let’s do now the same task with rray.
library(rray)
(mat_1_rray <- rray(c(15, 10, 8, 6, 12, 9), dim = c(2, 3)))
(mat_2_rray <- rray(c(5, 2, 3), dim = c(1, 3)))
## Broadcasting works ✓
mat_1_rray + mat_2_rray
## Subsetting works ✓
dim(mat_1_rray[,2:3])
dim(mat_1_rray[,1])
## smart functions
mat_1_rray / rray_sum(mat_1_rray, axes = 1)
rray_bind(mat_1_rray, mat_2_rray, .axis = 1)
rray_bind(mat_1_rray, mat_2_rray, .axis = 2)More info
- Program + Slides: https://user2019.r-project.org/talk_schedule/
- Collection of slides during the conference by Praer (Suthira Owlarn): https://github.com/sowla/useR2019-materials
- Recordings by the R Consortium on Youtube (keynotes available, rest soon to be published)
Last but not least
The rstatsmeme package is a little gem discovered thanks to Frie Preu:
# devtools::install_github("favstats/rstatsmemes")
library(rstatsmemes)
show_me_an_R_meme()
I cannot wait to see the recordings to catch up with the parallel sessions that I missed!↩
I can totally confirm that.↩
If you can, choose the UKBB + GIANT meta analysis results, which are pretty large.↩
Apparently, characters are the most challenging ones for speed.↩